Welcome to BEELINE’s documentation!
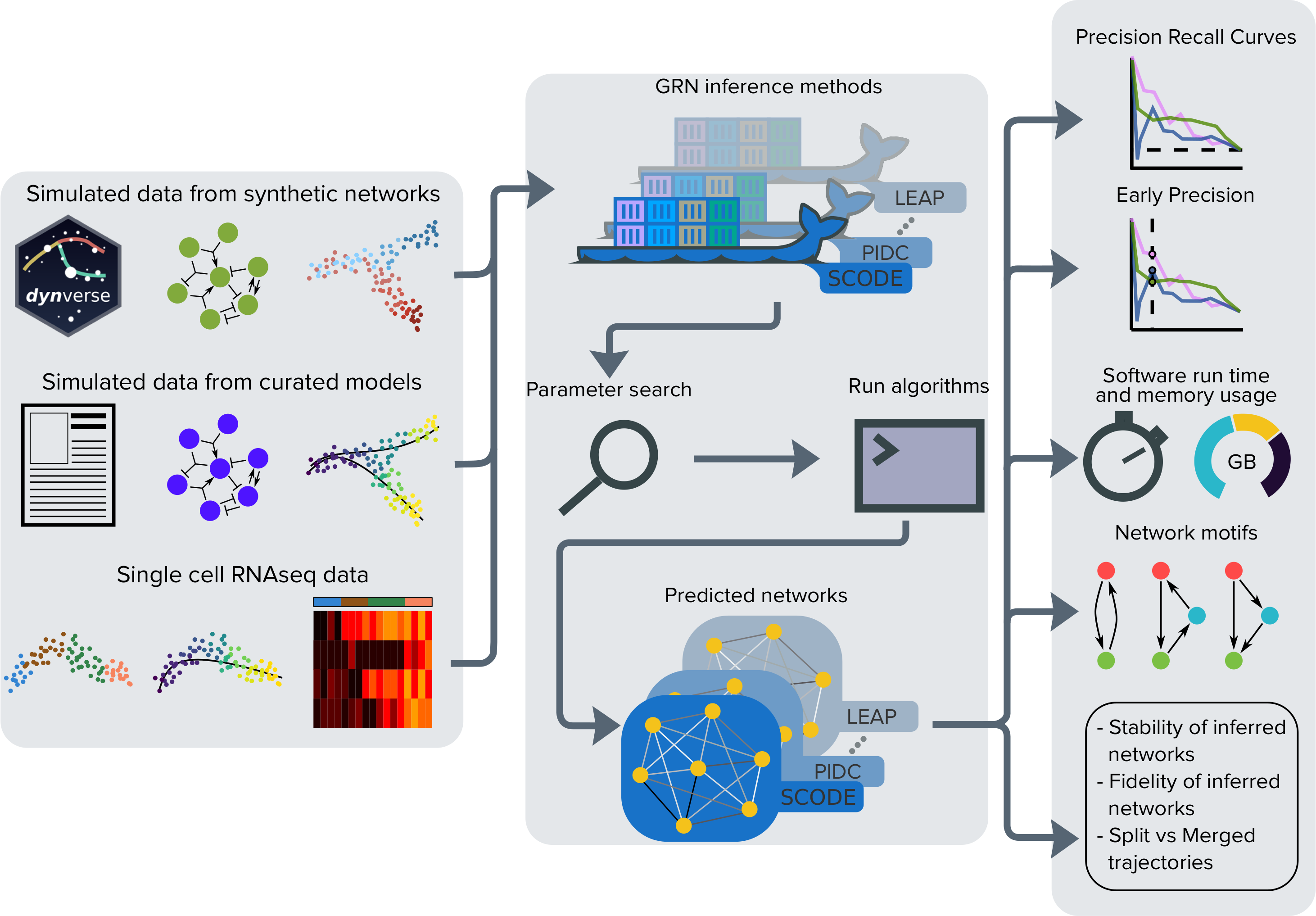
Overview of BEELINE
This site serves as the documentation for the code released under the Benchmarking gEnE reguLatory network Inference from siNgle-cEll transcriptomic data (BEELINE) project. The first part of this project is the BEELINE pipeline which provides tools to evaluate the performance of algorithms for the reconstruction of gene regulatory networks (GRNs) from single-cell RNAseq data. The second part of this project is BoolODE, a tool to automatically convert a Boolean model to an ODE model, and subsequently carry out stochastic simulations.
BEELINE provides a set of tools for evaluating methods that infer gene regulatory networks (GRN) from single-cell gene expression data. The BEELINE framework is divided into the following modules:
BLRun package : contains BEELINE’s Runner module, a Python wrapper for 14 GRN inference algorithms with options to add new methods.
BLEval package : contains BEELINE’s Evaluation module that provides easy-to-use tools for evaluating GRN reconstructions.
BLPlot package : contains BEELINE’s plotting module for visualizing output from BLEval.